Python Example - CFEM
Navigation: User Guide ➔ COM Automation ➔ Python Automation ➔ Example - CFEM
| Python Setup | Python Examples | Python Script | Optimisation | Visulisation | Python GUI | Tags and Data | |||
|---|---|---|---|---|---|---|---|---|---|
| Installation & Troubleshooting | List of Examples | Simple Script (pywin32) | SysCAD COM Python Class | Automated Model Testing | Constrained FEM (numpy|matplotlib) | Optimisation (numpy|scipy) | Adding Plots (numpy|matplotlib) | Dynamic with GUI | Accessing Data (sqlite3|pandas) |
Constrained Free Energy Minimization
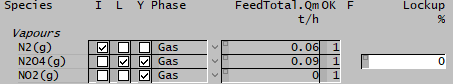
| Python’s numerical libraries are highly effective for manipulating large data arrays and are essential for performing complex calculations. This example demonstrates time evolution of [math]\ce{ N2O4 }[/math] decomposition using GFEM lockup to constrain the reaction [math]\ce{ N2O4 = 2 NO2 }[/math] and then calculate the evolution of the reaction. We conducted a series of GFEM equilibrium simulations involving the species N₂O₄, varying its constraint from fully locked (100%) to completely unconstrained (0%, representing equilibrium conditions). SysCAD project Configuration information:
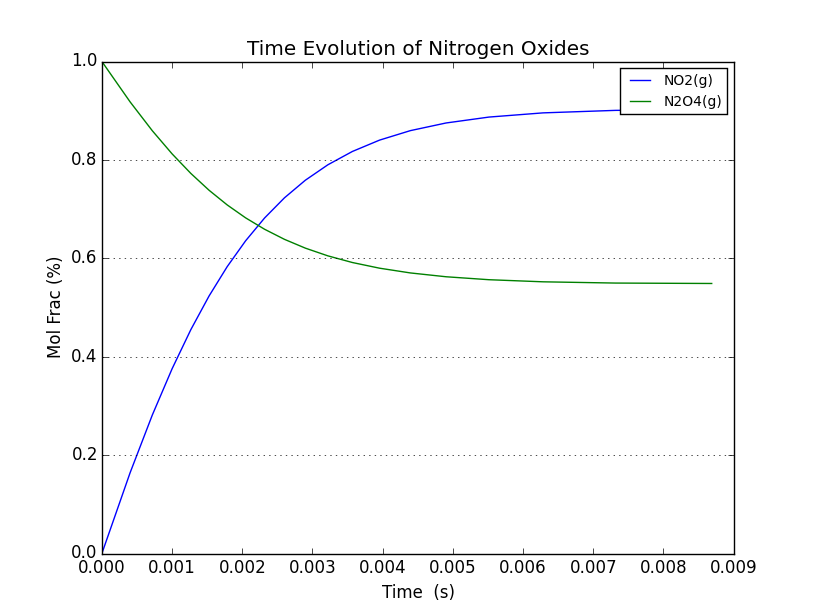
Using the resulting data, we applied a rate equation to calculate the time required for each incremental change and plotted the time evolution of the system. The rate equation is 1
Rather than integrate this directly, we rewrite as:
which determines the time [math]\displaystyle{ \Delta t }[/math] for a change in concentration. Reference
|
import time
import math
import matplotlib.pyplot as plt ## Plotting graphics
import numpy as np ## Numerical python
import syscadif ## SysCAD COM interface class
sc = syscadif.SysCADCom()
ScdDir = r'C:\SysCAD%d_37043' % sc.build ## Installation folder
ScdPrj = r'\Examples\03 UnitModels\GFEM Simple Examples.spf\Project.spj'
sc.OpenProject(ScdDir+ScdPrj) # Open the SysCAD project
Tags = sc.Tags
ProBal = sc.Solver.ProBal
Prod = [
"NO2(g)" , # 0
"N2O4(g)" , # 1
]
prodTagLis = [("P_002.Qo.QMl.%s (kmol/h)" % c) for c in Prod]
l1 = 100*(1.-np.linspace(0, 1, 20)**2) ## generate a linear space but increase resolution by squaring it
lkup = l1[::-1] ## reverse it
def genData():
sc.setTags(["FEMR_001.T_Reqd (C)"], [40])
sc.setTags(["FEMR_001.OperatingP.P_Reqd (atm)"], [1.0])
res = [(0.0, 1.0)]
res.extend(sc.RunScenarios(100-lkup[1:], "FEMR_001.FEMD.[N2O4(g)].Lockup (%)", prodTagLis))
return np.array(res)
rp = genData()
Ea = 40.e3 ##81.e3
R = 8.3144
T_K = 273.15+40
A = 1.0e9
k = A*math.exp(-Ea/(R*T_K))
rr = rp/rp[-1] ## Concentration/EquilibriumConcentration
Affinity = 1.-rr[:, 0]**2/rr[:, 1] ## Affinity
DeltaN2O4 = -np.diff(rp[:, 1]) ## Change in [N2O4]
N2O4Term = k*(rp[:-1, 1]+rp[1:, 1])/2. ## Average value of [N2O4] in each time step
DeltaT = DeltaN2O4/N2O4Term/Affinity[:-1] ## Delta Time for each reaction extent change
T = np.insert(np.cumsum(DeltaT), 0, 0) ## Actual (cumulative) time
def doPlot():
lockup_values = 100 - lkup # Lockup (%) values
fig, axs = plt.subplots(2, 1, figsize=(10, 8), sharex=True)
# First subplot: NO2 and N2O4
for Y, lbl in zip(rp.transpose(), ["NO2(g) Produced", "N2O4(g) Remaining"]):
axs[0].plot(T, Y, label=lbl, marker='x')
# Annotate Lockup (%) values at selected points
for i in range(0, len(T), 19): # every 19 point
axs[0].annotate("at equilibrium", (T[-1], rp[-1, 1]),
textcoords="offset points", xytext=(0,5),
ha='center', fontsize=8, color='purple')
axs[0].legend(fontsize='small', loc='lower right')
axs[0].set_ylabel("Flowrate (kmol/h)")
axs[0].grid(True, linestyle="-", color='lightgrey')
axs[0].set_title("Time Evolution of Nitrogen Oxides")
# Second subplot: Lockup (%)
axs[1].plot(T, lockup_values, label='Lockup (%)', linestyle='--', color='purple', marker='o')
# Annotate Lockup (%) values at selected points
for i in range(0, len(T), 2): # every 3rd point
axs[1].annotate(f"{lockup_values[i]:.1f}%", (T[i], lockup_values[i]),
textcoords="offset points", xytext=(5,0),
ha='left', fontsize=8, color='purple')
axs[1].set_ylabel("Lockup (%)")
axs[1].set_xlabel("Time (s)")
axs[1].grid(True, linestyle="-", color='lightgrey')
axs[1].set_title("N2O4 Lockup Percentage")
axs[1].legend(fontsize='small', loc='upper right')
plt.show()
input('Enter to finish')
sc.CloseProject()
sc.Close()
doPlot()
|